Research Program 2
Metabolic Dynamics and Organ Crosstalk
The goal of this Research Program is to uncover the stream of molecules that are released from cells in response to energetic stressors, and to understand what regulates these processes during healthy and cardiometabolic disease conditions.
Scope
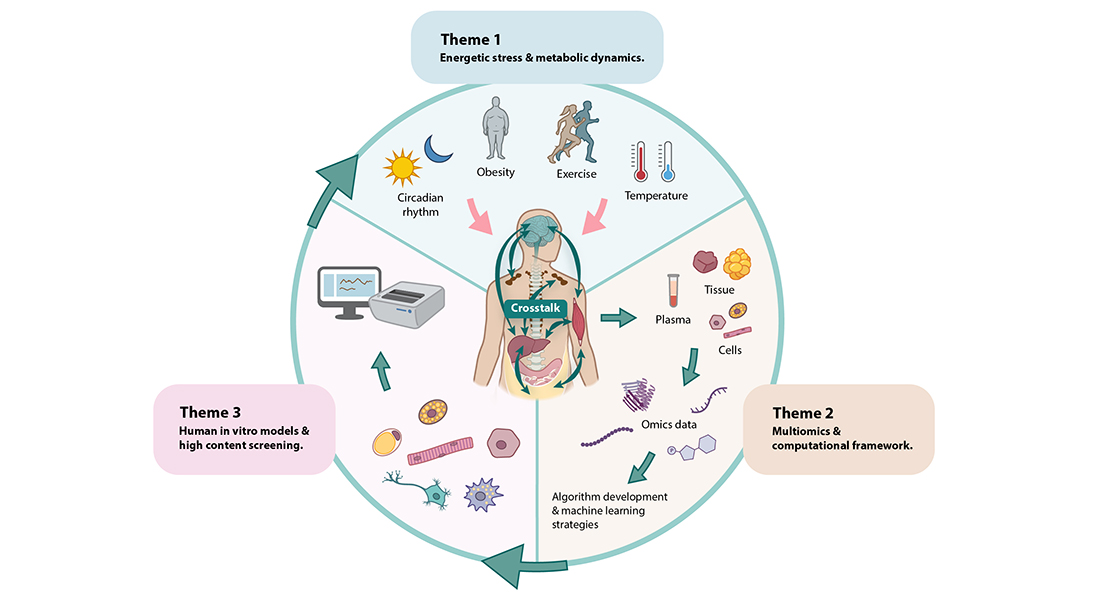
Program 2 also aims to understand processes that are sensitive to, or could affect, the response to energetic stressors. These processes include circadian rhythm, stellate cell activation, myogenesis, adipogenesis, insulin sensitivity and NAD+ metabolism. The science in Program 2 builds on the perception of an efficient communication of energetic states between organs and cells as a prerequisite for maintaining cardiometabolic health in humans (Figure 15). Ultimately, Program 2 asks: what are the pathophysiological events leading to a dysfunctional organ crosstalk, and how can this be reversed?
Program 2 will harness multidisciplinary approaches including cutting-edge omics technologies, human clinical samples, and human in vitro models for studies of dynamic responses to energetic stressors, as well as high-content screening for functional validation of computationally-prioritized candidate signaling molecules. Program 2 will focus on metabolically important organs such as liver, skeletal muscle and adipose tissue and their responses to energetic stressors. Program 2 will provide deep insight into the metabolic organ crosstalk events and identify new drug-target candidates for de-risking prevention and treatment of cardiometabolic diseases.
The scope of Program 2 does not extend to research on kidney function and secretome in cardiometabolic disease, or to research on type 1 diabetes.
The focus of this research theme is to understand how energetic stress, including exercise and temperature, interacts with circadian rhythm to regulate a healthy metabolism and counteract obesity and cardiometabolic disease. Despite extensive research, the molecular mechanisms behind the protective effects of exercise remain incompletely understood. A major challenge in the field is the complexity of the effectors, the multi-organ involvement, and the dynamic signaling network (e.g., via multiple post-translational modifications). To address this challenge, Program 2 will combine a multi-omics approach with clinical samples and robust phenotyping of genetically modified mouse models. Program 2 will collaborate with the Data Analytics Platform to develop computational strategies for joint modeling of exercise-induced secreted proteins, peptides, and metabolites to capture the global effects of exercise and to understand how specific organs communicate to facilitate appropriate adaptations.
The Program will apply in vitro and in vivo expertise in generating adeno-associated viruses and CRISPR-mediated gene editing in cell and animal models to manipulate the identified signaling factors and assess downstream effects. Pharmacological studies will be performed in collaboration with the Rodent Metabolic Phenotyping Platform and Research Program 3. Ultimately, this research theme aims to provide deeper insight in molecular circuits and pathways that affect the pathophysiology of insulin resistance and might help restore metabolic health.
Understanding the complex metabolic crosstalk and how this is dysregulated during cardiometabolic disease requires sampling and analyses across multiple tissues and conditions. Program 2 has a record of accomplishment in performing cutting-edge omics studies addressing the effects of energetic stress (PMIDs: 33436631, 31668873, 35030324, 34038727, 35638262). Program 2 will now take the leap into application of multiomics technologies across tissue types, cell types and body fluid samples, harvested during distinct metabolic conditions including exercise, cold exposure, obesity, nonalcoholic steatohepatitis and cardiometabolic disease.
To better understand the organ crosstalk network during or in response to these conditions, a major part of this research theme will be to develop a computational framework for joint modeling of omics datasets and for developing machine learning based software for candidate prioritization. Organ crosstalk is a major focus and Program 2 will thus map secreted proteins, peptides, microproteins and metabolites. This will be achieved by performing proteomics, extracellular vesicle proteomics, peptidomics, metabolomics, full-length RNA sequencing, ribosomal sequencing as well as single-cell proteomics and single-cell and single-nuclei RNA sequencing. Assembling these data sets will also allow for connecting secretome data with intra-organ and intra-cellular data and developing new hypotheses.
To test specific hypotheses generated within Theme 1 or 2, and/or to perform unbiased screening, Program 2 will develop advanced human in vitro models including 3D-cultures, multicellular co-culture systems, and organoids, using human primary cultures and hiPSC. Initially, Program 2 will focus on modeling adipose tissue, liver, skeletal muscle and neurons. However, this could be expandable to other tissues should it become relevant. The in vitro models will be examined by single-cell transcriptomics and will be applied in high-content screening systems including automated imaging-based assays and multi-well electrophysiology. The Center has recently invested in a Leica Thunder Imager Live Cell and 3D assay microscope as well as a Maestro Pro multiwell microelectrode array system, for recording high-content dynamic responses. Furthermore, Center scientists have access to an ImageXpress Pico Automated Cell Imager system for fluorescence based high-content screening.
Program 2 works in close collaboration with clinical partners to access human material for isolation of progenitor cells from skeletal muscle and adipose tissue, as well as primary cultures of hepatocytes. The work will be performed in collaboration with the Single-Cell Omics Platform, the Data Analytics Platform, the Novo Nordisk Foundation Center for Protein Research and the Novo Nordisk Foundation Center for Stem Cell Medicine (reNEW).


